AI Tech Accurately Diagnoses Knee Arthritis from Medical Images
Vagheesh Narasimhan and Prakash Jayakumar trained an AI on x-ray images from tens of thousands of people in the UK Biobank.
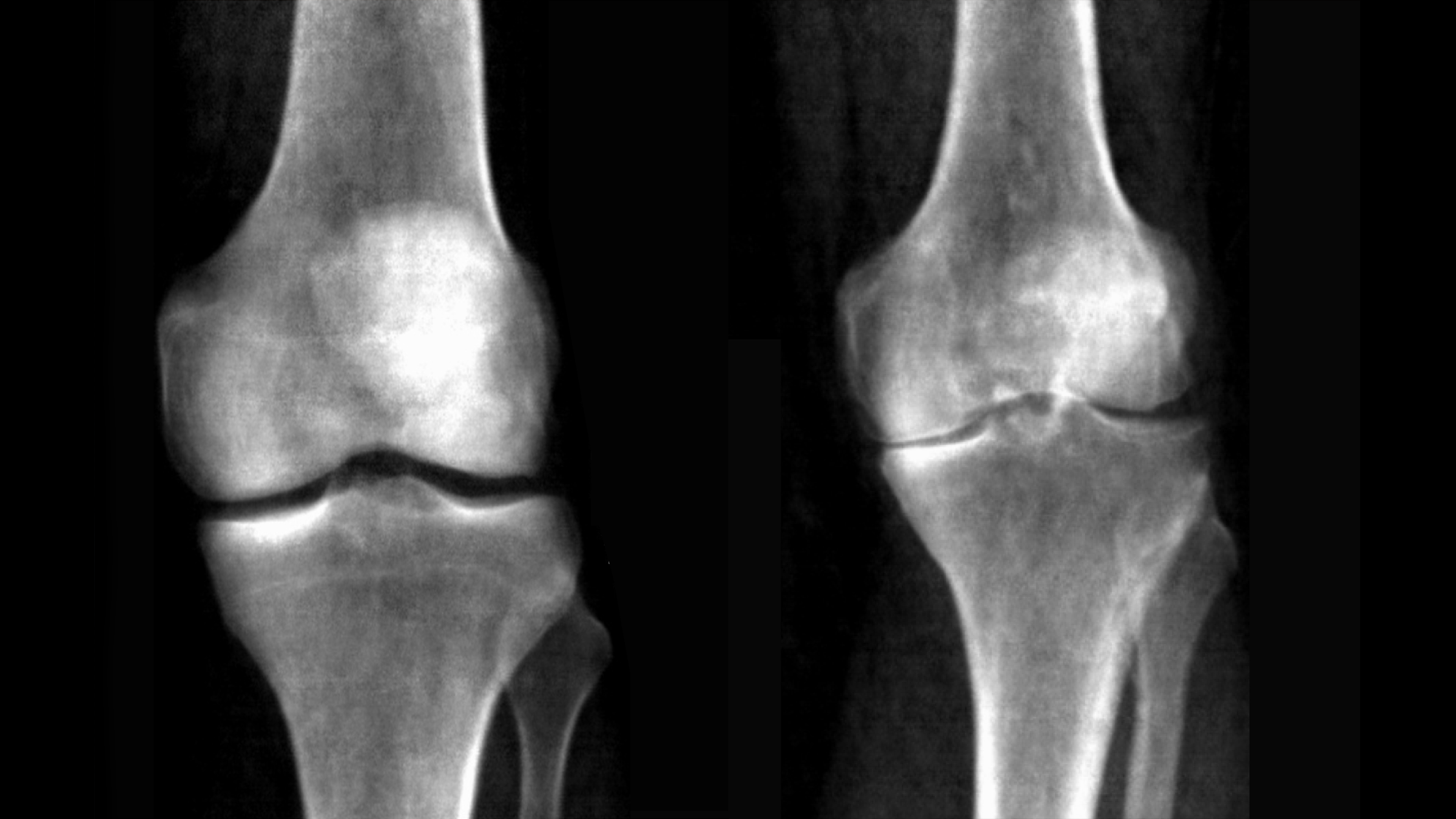
X-ray images of a healthy knee (left) compared to an individual with knee osteoarthritis. Credit: UK Biobank/University of Texas at Austin.
Many people over the age of 50 develop arthritis of the knee, a painful condition that can make it difficult to do everyday activities like stand, walk or climb stairs. New research published in Nature Digital Medicine on the use of artificial intelligence to analyze medical images demonstrates the potential for improving diagnoses of knee osteoarthritis and predicting a person’s risk of developing it in the future.
A team led by researchers from The University of Texas at Austin’s College of Natural Sciences and Dell Medical School developed an AI model that can diagnose knee osteoarthritis with clinical-grade performance, based solely on images of the knee joint from medical x-rays. One potential application would be incorporating the model into existing software that technicians use to evaluate x-rays, flagging possible cases of knee osteoarthritis to assist clinicians in making a diagnosis. The goal wouldn’t be to replace human doctors but to assist them.
“Imagine someone goes into the clinic and gets an x-ray for some other reason, and the clinic is running this AI system in the background on every image that automatically diagnoses arthritis,” said Vagheesh Narasimhan, an assistant professor in UT Austin’s Department of Statistics and Data Sciences and Department of Integrative Biology. “We could tell them, perhaps you should consult an orthopedic expert about this condition.”
The study’s co-first authors are Brianna Flynn and Emily Javan, graduate students in UT Austin’s Department of Integrative Biology. Co-senior authors are Narasimhan and Prakash Jayakumar, assistant professor in the Department of Surgery and Perioperative Care at UT Austin’s Dell Medical School.
Because arthritis of the knee is caused by the grinding of knee bones (the femur and tibia) against each other as protective cartilage wears down, the size of the gap between the bones is associated with how severe the condition is. This gap, known as the joint space can be used as a clinical biomarker to quantitatively assess disease severity. In their work the researchers were also able to provide automatic measurements on the knee joint space with high precision and replicability. This quantitative measure could help doctors more accurately track disease progression in one person over time.
Arthritis of the knee is a leading cause of adult disability in the U.S., and it’s very costly to intervene once it develops into a severe case.
“So we’re also thinking about ways to combine imaging, genetic data and other risk factors simultaneously into a single model, to provide a more comprehensive prediction for knee osteoarthritis,” Narasimhan said.
The team trained their AI model on medical records from tens of thousands of people in the UK Biobank, a large-scale biomedical database and research resource, containing in-depth genetic and health information from half a million UK participants.
Using their model, the researchers diagnosed nearly twice as many cases of knee arthritis as were documented in the UK Biobank health records. There are many possible reasons for the unreported cases in the medical records. Clinicians might set arbitrary thresholds, choosing not to diagnose milder cases. Patients themselves might ignore or downplay the condition due to varying levels of pain tolerance, or because they don’t want to consider treatment options such as surgery due to the high cost and long recovery time.
Filling in these gaps could help researchers sifting through large sets of health records in search of factors that might correlate with disease such as genetics, lifestyle (e.g., nutrition, exercise, addiction or medications) and environmental factors (e.g., pollutants or accidents).
“An unbiased means of performing diagnosis on the entire cohort of half a million people is kind of exciting and will help improve those types of epidemiological association studies,” Narasimhan said.
Other UT Austin authors are: Eugenia Lin, Zoe Trutner, Karl Koenig, Kenoma Anighoro, Eucharist Kun and Alaukik Gupta. The study’s other author is Tarjinder Singh, an assistant professor in Columbia University’s Department of Psychiatry, with joint appointments at the Mortimer B. Zuckerman Mind Brain Behavior Institute and the New York Genome Center.
Many of the same collaborators published another paper earlier this summer in Science applying the same AI system and dataset to identify genetic markers related to skeletal health and development.
This research was supported in part by a Good Systems for Ethical AI grant from UT Austin. Additional support was provided by the Paul G. Allen Family Foundation, the National Science Foundation and the National Institutes of Health.



